|
|
|
|
|
| Searching in 'World-2DPAGE Repository [0069]' for entry matching: DHSA_RAT |
World-2DPAGE Repository (0069): DHSA_RAT
DHSA_RAT
| General information about the entry |
| View entry in simple text format |
| Entry name | DHSA_RAT |
| Primary accession number | DHSA_RAT |
| integrated into World-2DPAGE Repository (0069) on | February 27, 2014 (release 1) |
| 2D Annotations were last modified on | February 27, 2014 (version 1) |
| General Annotations were last modified on | February 27, 2014 (version 1) |
| Name and origin of the protein |
| Description | DHSA_RAT. |
| Annotated species | Rattus [TaxID: 10114] |
| Taxonomy | Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Glires; Rodentia; Sciurognathi; Muroidea; Muridae; Murinae. |
| References |
| [1] |
2D GEL CHARACTERIZATION
DOI=10.1016/j.jprot.2014.04.015;
Burniston J.G., Kenyani J., Gray D., Guadagnin E., Jarman I.H., Cobley J.N., Cuthbertson D.J., Chen Y.-W., Wastling J.M., Lisboa P., Koch L.G., Britton S.L.
''Conditional independence mapping of DIGE data reveals PDIA3 protein species as key nodes associated with muscle aerobic capacity''
Journal of Proteomics 106(1):230-245 (2014)
|
| | 2D PAGE maps for identified proteins
|
|
How to interpret a protein
|
RATTUS_SOLEUS-DIGE_4.5-9.5 {M+F HCR LCR Soleus DiGE Exp}
Rattus
Tissue: Skeletal muscle
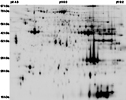
map experimental info
|
|
RATTUS_SOLEUS-DIGE_4.5-9.5
MAP LOCATIONS:
% COVERAGE: SPOT 42: 19 [1]
SPOT 57: 9 [1]; SPOT 87: 63 [1]; SPOT 91: 35 [1]; SPOT 92: 38 [1]; SPOT 95: 58 [1]; SPOT 127: 33 [1]; SPOT 138: 45 [1]; SPOT 155: 5 [1].
MOWSE: SPOT 42: 522 [1]
SPOT 57: 123 [1]; SPOT 87: 1649 [1]; SPOT 91: 924 [1]; SPOT 92: 1163 [1]; SPOT 95: 1625 [1]; SPOT 127: 1105 [1]; SPOT 138: 1232 [1]; SPOT 155: 145 [1].
NAME: SPOT 42: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]
SPOT 57: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 87: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 91: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 92: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 95: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 127: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 138: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1]; SPOT 155: Succinate dehydrogenase [ubiquinone] flavoprotein subunit, m [1].
| | Copyright |
| Data from Dr. Jatin G. Burniston, Research Institute for Sport and Exercise Sciences, Liverpool John Moores University, UK |
World-2DPAGE Repository (search AC)
|