| General information about the entry |
| View entry in simple text format |
| Entry name | ODPB_MOUSE |
| Primary accession number | Q9D051 |
| integrated into World-2DPAGE Repository (0027) on | September 30, 2010 (release 1) |
| 2D Annotations were last modified on | June 21, 2011 (version 2) |
| General Annotations were last modified on | November 23, 2011 (version 2) |
| Name and origin of the protein |
| Description | RecName: Full=Pyruvate dehydrogenase E1 component subunit beta, mitochondrial; Short=PDHE1-B; EC=1.2.4.1; Flags: Precursor;. |
| Gene name | Name=Pdhb
|
| Annotated species | Mus musculus (Mouse) [TaxID: 10090] |
| Taxonomy | Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Glires; Rodentia; Sciurognathi; Muroidea; Muridae; Murinae; Mus; Mus. |
| References |
| [1] |
2D-PAGE GEL CHARACTERIZATION
DOI=10.1002/prca.201000103;
Blutke A., Block C., Berendt F., Herbach N., Kemter E., Amann K., Frolich T., Arnold G.J., Wanke R.
''Differential glomerular proteome analysis of two murine nephropathy models at onset of albuminuria''
Proteomics - Clinical Applications 5(6):375-381 (2011)
|
|
| 2D PAGE maps for identified proteins
|
|
How to interpret a protein
|
GIPRDN_4-7 {glomerular protein lysates of a GIPRdn-tg animal}
Mus musculus (Mouse)
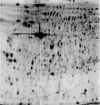
map experimental info
|
|
GIPRDN_4-7
MAP LOCATIONS:
EXPRESSION:
| SPOT 25: Avg ratio GIPRdn-tg vs. control=1.84 (p-value=0.0064) [1]. |
IDENTIFICATION: SPOT 25: SeqCov=8%. Mascot protein score=62 [1].
MAPPING (identification):
|
| Copyright |
| Data from Dr. Andreas Blutke, Institute of Veterinary Pathology, University Munich, Germany |
| Cross-references |
| REPRODUCTION-2DPAGE | Q9D051; Q9D051. |
| UCD-2DPAGE | Q9D051; ODPB_MOUSE. |
| UniProtKB/Swiss-Prot | Q9D051; ODPB_MOUSE. |
| World-2DPAGE Repository | Q9D051; ODPB_MOUSE. |