Attention: World-2DPAGE is no longer maintained.
[Click to access protein entry]
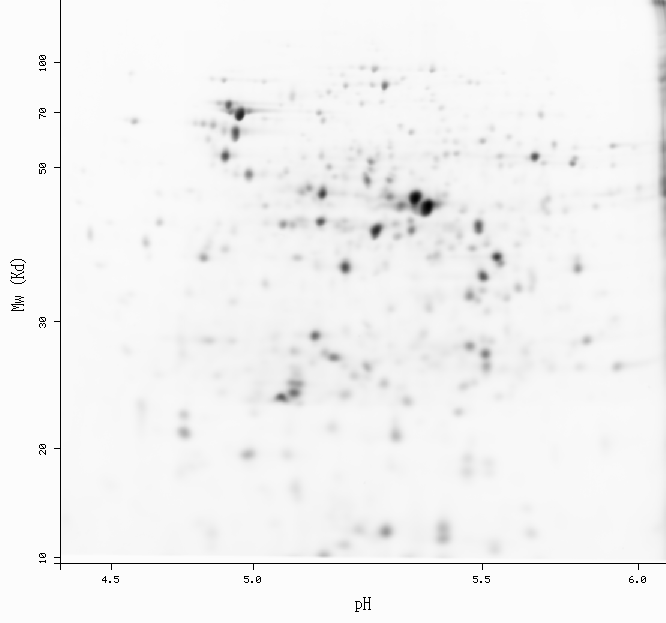
Spot: 2D-001WM0 (ecoli-dige4.5-6.5)
pI: 5.88 Mw: 42128
%vol: 0.133919 %od: 0.105318
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS} {PMF}
peptide sequences: {
$m/Z=(K)SALLVLEDGTQFHGR(A);; $m/Z=(R)NTEDLSSYLK(R);; $m/Z=(K)EVTTAEAYSWTQGSWTLTGGLPEAK(K);; $m/Z=(K)KEDELPFHVVAYDFGAK(R);; $m/Z=(R)LTIVPAQTSAEDVLK(M);; $m/Z=(K)FLETDIPVFGICLGHQLLALASGAK(T);; $m/Z=(K)SLFDGTLQGIHR(T) }
peptide masses: { (TRYPSIN)
$m/Z= 1145.51 (0), 1186.47 (0), 1325.55 (0), 1337.59 (0), 1343.59 (0),
1493.66 (0), 1642.72 (0), 1657.65 (0), 1964.75 (0), 4064.65 (0)
}
Spot: 2D-001WM0 (ecoli-dige4.5-6.5)
pI: 5.88 Mw: 42128
%vol: 0.133919 %od: 0.105318
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS} {PMF}
peptide sequences: {
$m/Z=(K)SALLVLEDGTQFHGR(A);; $m/Z=(R)NTEDLSSYLK(R);; $m/Z=(K)EVTTAEAYSWTQGSWTLTGGLPEAK(K);; $m/Z=(K)KEDELPFHVVAYDFGAK(R);; $m/Z=(R)LTIVPAQTSAEDVLK(M);; $m/Z=(K)FLETDIPVFGICLGHQLLALASGAK(T);; $m/Z=(K)SLFDGTLQGIHR(T) }
peptide masses: { (TRYPSIN)
$m/Z= 1145.51 (0), 1186.47 (0), 1325.55 (0), 1337.59 (0), 1343.59 (0),
1493.66 (0), 1642.72 (0), 1657.65 (0), 1964.75 (0), 4064.65 (0)
}
[Click to access protein entry]
Spot: 2D-001WMA (ecoli-dige4.5-6.5)
pI: 5.79 Mw: 41417
%vol: 0.115129 %od: 0.102482
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)DLPLIASNFR(N);; $m/Z=(R)LTIVPAQTSAEDVLK(M);; $m/Z=(K)FLETDIPVFGICLGHQLLALASGAK(T);; $m/Z=(K)SLFDGTLQGIHR(T) }
Spot: 2D-001WMA (ecoli-dige4.5-6.5)
pI: 5.79 Mw: 41417
%vol: 0.115129 %od: 0.102482
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)DLPLIASNFR(N);; $m/Z=(R)LTIVPAQTSAEDVLK(M);; $m/Z=(K)FLETDIPVFGICLGHQLLALASGAK(T);; $m/Z=(K)SLFDGTLQGIHR(T) }
[Click to access protein entry]
Spot: 2D-001WM3 (ecoli-dige4.5-6.5)
pI: 5.82 Mw: 42128
%vol: 0.0512572 %od: 0.0314685
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)DLPLIASNFR(N);; $m/Z=(R)NTEDLSSYLKR(H) }
Spot: 2D-001WM3 (ecoli-dige4.5-6.5)
pI: 5.82 Mw: 42128
%vol: 0.0512572 %od: 0.0314685
================
*CARA_ECOLI*
accession n°: P0A6F1
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)DLPLIASNFR(N);; $m/Z=(R)NTEDLSSYLKR(H) }
SWISS-2DPAGE Viewer
|
Back to the |
-Get more information by dragging your mouse pointer over any highlighted spot -Click on a highlighted spot to access all its associated protein entries
[estimated location]
Back to the