Attention: World-2DPAGE is no longer maintained.
[Click to access protein entry]
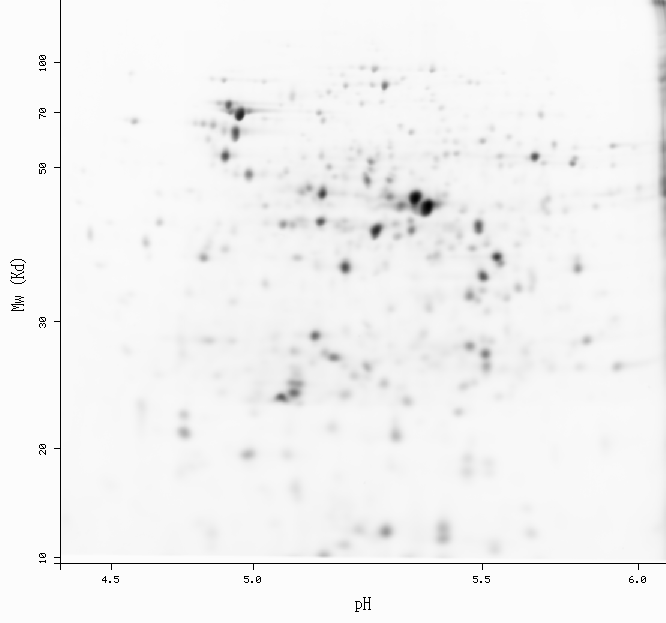
Spot: 2D-001WI2 (ecoli-dige4.5-6.5)
pI: 5.15 Mw: 52681
%vol: 0.187731 %od: 0.161483
================
*GPMI_ECOLI*
accession n°: P37689
Identification Methods:
* MAPPING: {MS/MS} {PMF}
peptide sequences: {
$m/Z=(R)AFFANPVLTGAVDK(A);; $m/Z=(K)AYDLLTLAQGEFQADTAVAGLQAAYAR;; $m/Z=(R)DENDEFVK(A);; $m/Z=(R)AFVNADFDGFAR(K);; $m/Z=(K)YAHVTFFFNGGVEESFKGEDR(I) }
peptide masses: { (TRYPSIN)
$m/Z= 861.35 (0), 874.42 (0), 987.46 (0), 995.32 (0), 1007.46 (0),
1204.51 (0), 1329.47 (0), 1449.56 (0), 1569.57 (0), 1625.63 (0),
1770.7 (0), 1978.64 (0), 2435.82 (0), 2514.98 (0), 2827.06 (0),
2984.07 (0), 3026.98 (0), 3043.99 (0) }
Spot: 2D-001WI2 (ecoli-dige4.5-6.5)
pI: 5.15 Mw: 52681
%vol: 0.187731 %od: 0.161483
================
*GPMI_ECOLI*
accession n°: P37689
Identification Methods:
* MAPPING: {MS/MS} {PMF}
peptide sequences: {
$m/Z=(R)AFFANPVLTGAVDK(A);; $m/Z=(K)AYDLLTLAQGEFQADTAVAGLQAAYAR;; $m/Z=(R)DENDEFVK(A);; $m/Z=(R)AFVNADFDGFAR(K);; $m/Z=(K)YAHVTFFFNGGVEESFKGEDR(I) }
peptide masses: { (TRYPSIN)
$m/Z= 861.35 (0), 874.42 (0), 987.46 (0), 995.32 (0), 1007.46 (0),
1204.51 (0), 1329.47 (0), 1449.56 (0), 1569.57 (0), 1625.63 (0),
1770.7 (0), 1978.64 (0), 2435.82 (0), 2514.98 (0), 2827.06 (0),
2984.07 (0), 3026.98 (0), 3043.99 (0) }
[Click to access protein entry]
Spot: 2D-001WIC (ecoli-dige4.5-6.5)
pI: 5.11 Mw: 52363
%vol: 0.119547 %od: 0.0810386
================
*GPMI_ECOLI*
accession n°: P37689
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)EEQQDNAIFSAK(T);; $m/Z=(R)AFFANPVLTGAVDK(A);; $m/Z=(K)AYDLLTLAQGEFQADTAVAGLQAAYAR;; $m/Z=(R)AFVNADFDGFAR(K) }
Spot: 2D-001WIC (ecoli-dige4.5-6.5)
pI: 5.11 Mw: 52363
%vol: 0.119547 %od: 0.0810386
================
*GPMI_ECOLI*
accession n°: P37689
Identification Methods:
* MAPPING: {MS/MS}
peptide sequences: {
$m/Z=(R)EEQQDNAIFSAK(T);; $m/Z=(R)AFFANPVLTGAVDK(A);; $m/Z=(K)AYDLLTLAQGEFQADTAVAGLQAAYAR;; $m/Z=(R)AFVNADFDGFAR(K) }
SWISS-2DPAGE Viewer
|
Back to the |
-Get more information by dragging your mouse pointer over any highlighted spot -Click on a highlighted spot to access all its associated protein entries
[estimated location]
Back to the