| General information about the entry |
| View entry in simple text format |
| Entry name | RLP24_HUMAN |
| Primary accession number | Q9UHA3 |
| integrated into SWISS-2DPAGE on | May 15, 2003 (release 16) |
| 2D Annotations were last modified on | December 30, 2004 (version 1) |
| General Annotations were last modified on | May 19, 2011 (version 6) |
| Name and origin of the protein |
| Description | RecName: Full=Probable ribosome biogenesis protein RLP24; AltName: Full=Ribosomal L24 domain-containing protein 1; AltName: Full=Ribosomal protein L24-like;. |
| Gene name | Name=RSL24D1
Synonyms=C15orf15, RPL24L
ORFNames=My024
|
| Annotated species | Homo sapiens (Human) [TaxID: 9606] |
| Taxonomy | Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Primates; Haplorrhini; Catarrhini; Hominidae; Homo. |
| References |
| [1] |
MAPPING ON GEL
PubMed=12429849; [NCBI, Expasy, EBI, Israel, Japan]
Scherl A., Coute Y., Deon C., Calle A., Kindbeiter K., Sanchez J.-C., Greco A., Hochstrasser D.F., Diaz J.-J.
''''''Functional proteomic analysis of human nucleolus'';'';''
Mol. Biol. Cell. 13(1):4100-4109(2002)
|
|
| 2D PAGE maps for identified proteins
|
|
How to interpret a protein
|
NUCLEOLI_HELA_1D_HUMAN {SDS-PAGE of nucleolar proteins from Human HeLa cells}
Homo sapiens (Human)
Tissue: Cervix carcinoma
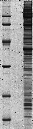
map experimental info
protein estimated location
|
|
NUCLEOLI_HELA_1D_HUMAN
MAP LOCATIONS:
MAPPING (identification):
Tandem mass spectrometry [1].
|
| Copyright |
| This SWISS-2DPAGE entry is copyright the Swiss Institute of Bioinformatics. There are no restrictions on its use by non-profit institutions as long as its content is in no way modified and this statement is not removed. Usage by and for commercial entities requires a license agreement (See http://world-2dpage.expasy.org/swiss-2dpage/docs/license.html or send email from legal@sib.swiss). |
| Cross-references |
| UniProtKB/Swiss-Prot | Q9UHA3; RLP24_HUMAN. |
| 2D PAGE maps for identified proteins
|
- How to interpret a protein map
- You may obtain an estimated location of the protein on various 2D PAGE maps, provided the whole amino acid sequence is known. The estimation is obtained according to the computed protein's pI and Mw.
- Warning 1: the displayed region reflects
an area around the theoretical pI and molecular weight of the protein and is only provided for the user's information.
It should be used with caution, as the experimental and theoretical positions of a protein may differ significantly.
- Warning 2: the 2D PAGE map is built on demand. This may take some few seconds to be computed.
|